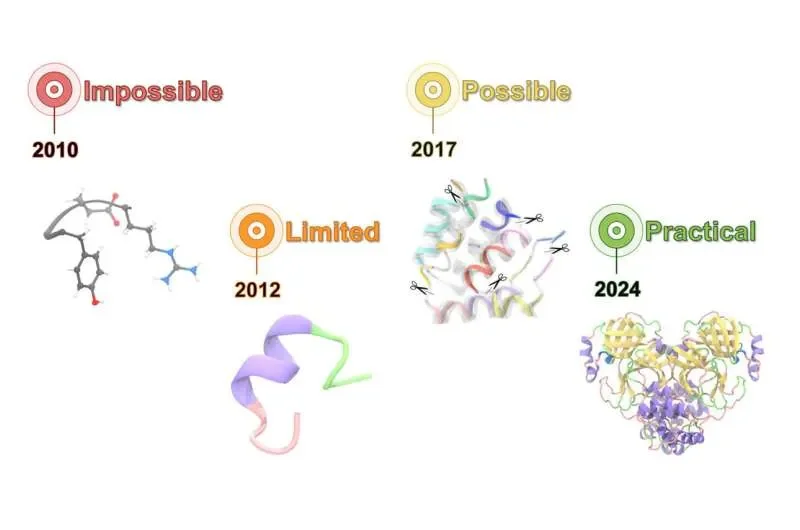
Berkeley Lab researchers have released AQuaRef, a calculation tool designed to fix the blurry maps of protein structures. This program merges quantum-mechanical math with machine learning to sharpen the jagged edges of molecular models. By shifting how computers guess where atoms sit, the tool cuts the high computational cost that usually stalls detailed biology.
The system moves away from crude approximations, instead using heavy math to predict how electrons and nuclei interact in both healthy and broken biological states.
The study, appearing in Nature Communications, points to a specific utility in metalloproteins—complex structures involving metal ions that often baffle standard mapping software. The logic is simple: use better math to require fewer guesses. The program aims for a higher level of precision than previous "unrestrained" refinement methods, specifically targeting the weird, irregular shapes that cause diseases.
The Mechanics of the Refinement
The AQuaRef system operates as a filter for raw data coming from cryo-electron microscopy and crystallography. While old tools often smudge the positions of atoms, this method uses a machine learning layer to accelerate the heavy lifting of quantum physics.
Read More: New Light Copper Catalysts Make Chemical Reactions Easier
It looks for the most stable energy states within the protein.
It corrects the placement of metal atoms which often look like noise in standard scans.
The "refinement" process acts as a digital tightening of the protein's actual physical limits.
| Feature | Standard Refinement | AQuaRef System |
|---|---|---|
| Math Basis | Classical physics / Balls-and-springs | Quantum-mechanical / ML loops |
| Speed | Variable (often slow for precision) | Accelerated through neural training |
| Target | General proteins | Difficult-to-map / Metalloproteins |
| Error Rate | Higher at atomic scales | Lower, closer to physical reality |
Structural Background
Biological mapping has long been a struggle between seeing a general shape and knowing exactly where the "fingers" of a protein are. Historically, researchers used programs like phenix.refine to guess these structures. However, these older tools often rely on simplified molecular models that ignore the messy, sub-atomic pull of electrons.
The introduction of AQuaRef represents a shift toward "real-space" refinement. Instead of a pretty picture, it provides a data-heavy coordinate map. By treating the protein as a series of quantum interactions rather than just a 3D object, the lab hopes to see how drugs might actually latch onto these surfaces in a diseased state. This is not about a "cure," but about making the invisible map a bit less blurry.
Read More: Kristin Smart Remains Not Found After New Search at Suspect's Mother's Home