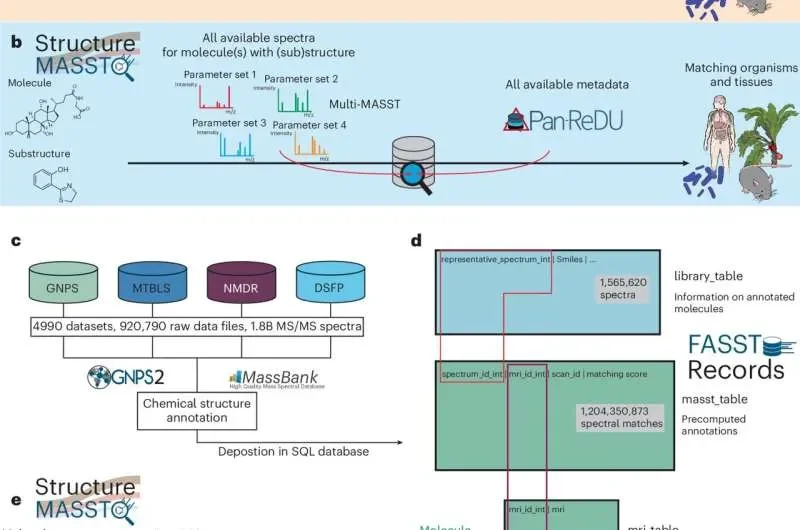
A novel search engine, StructureMASST, has been unveiled, aiming to untangle the complex web of life's chemistry. This tool aggregates data from major public metabolomics repositories, enabling researchers and the public alike to pinpoint molecules across diverse biological and environmental samples. The system allows for searches using chemical names, textual representations of molecular structures (SMILES strings), or partial molecular patterns, yielding instant results across various documented sources.
This groundbreaking search engine provides access to information concerning human, animal, plant, and even environmental samples. The documentation spans from the chemical signatures of recently extinct fauna to microbial communities found on the International Space Station. Search results can be further refined with tags indicating the organism, health status or disease, sample type, geographical location, and environmental context.
Bridging Data Silos, Demystifying Molecules
Historically, identifying specific molecules within public databases demanded specialized knowledge and was confined to disparate, isolated datasets. StructureMASST appears to dissolve these barriers by integrating a vast collection of information.
Read More: Red-footed Boobies Use Wind for Navigation, New Study Shows
"Until now, searching for specific molecules in public repositories required expert knowledge and was limited to isolated datasets."
The system's capacity to link molecular identity with contextual data – including organism origin, anatomical localization, associated health conditions, and environmental factors – marks a significant stride. For instance, a query for caffeine’s molecular structure reportedly returned over 6,000 matches, detailing its presence in coffee plants, human blood, breast milk, and cultured microbes.
Background: The Evolving Landscape of Molecular Inquiry
The development of StructureMASST signals a continuing trend towards making complex scientific data more accessible. Similar past endeavors, like the application of web search algorithms to analyze molecular interactions, underscore a persistent drive to translate raw data into understandable patterns.
The foundational research for StructureMASST has been detailed in a publication within Nature Biotechnology. The tool is a product of collaborative effort and technological innovation, potentially shaping future large-scale scientific investigations.