New algorithms are being used to map three-dimensional interactions within the cell nucleus, offering a novel way to understand how genes are switched on and off. Researchers at St. Jude Children's Research Hospital have developed an approach called BOUQUET, which leverages 'machine learning' to chart these complex cellular structures.
The core of this work lies in identifying "super-enhancers," regions of DNA that exert significant control over gene activity. Unlike simpler models that view DNA as a flat string, BOUQUET acknowledges the intricate, folded reality of the genome inside the nucleus. It suggests that these super-enhancers, along with the genes they influence, can cluster together in specialized compartments within the nucleus, referred to by researchers as "communities" or "protein condensates." These are described as dense, droplet-like structures without traditional membranes.
The Challenge of Distance and Dimension
Traditionally, understanding gene regulation has focused on a linear, two-dimensional view of DNA. However, the physical reality is a complex three-dimensional network. Enhancers, which act as switches for gene expression, can be located a considerable distance from the genes they control, sometimes separated by thousands of DNA bases. Mapping these distant connections in a 3D space presents a significant hurdle.
Read More: Space Weather May Block Alien Signals, Scientists Say
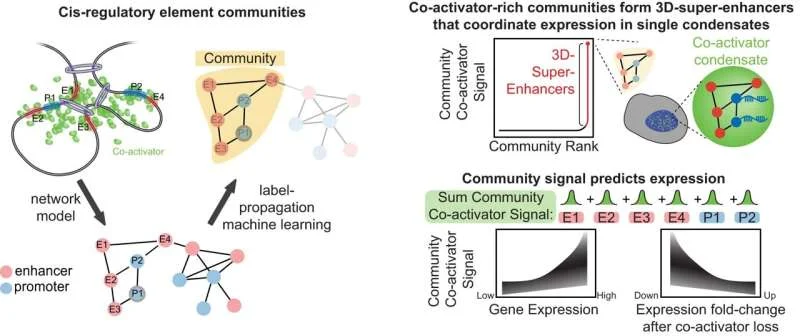
Algorithmic Insight into Cellular Organization
The development of BOUQUET signifies a shift towards understanding these interactions within their natural three-dimensional context. By training 'AI models' on various types of genomic data, including genetic sequences and multi-omic information, scientists aim to predict how different parts of the genome connect in 3D. This allows for the inference of "super-enhancer networks" and helps in pinpointing specific genetic variations that might alter how cells develop and change their fundamental identity.
Implications for Cell Fate and Future Study
The ability to map these 3D super-enhancers and their regulatory roles is becoming central to research on cell identity. Understanding these intricate connections could unlock new avenues for studying how cell fates are determined and potentially manipulated. For those looking to enter this field, developing skills in computational analysis and understanding of both genomics and cellular structures is highlighted as essential. The fundamental question remains: how does the folded, three-dimensional structure of the genome influence the activity of enhancers and, consequently, the defining characteristics of a cell?
Read More: Lorraine Kelly's 7-week absence from ITV show due to illness


