Digital Alchemists Generate Molecular Keys
The latest wave of computational tools, drawing heavily on AI-driven "diffusion models," is fundamentally reshaping how medicines are conceived. These systems construct entirely new drug molecules, aiming for precise fits with specific biological targets, a process that drastically shortens the timeline from initial idea to potential therapy.
This approach bypasses lengthy trial-and-error laboratory work. Instead, algorithms navigate vast digital landscapes of chemical possibilities. Researchers feed these systems information about a disease's protein targets, and the AI then designs molecules tailored to interact with them. This promises to accelerate the identification and refinement of new drug candidates, moving from conceptualization to evaluation at an unprecedented pace.
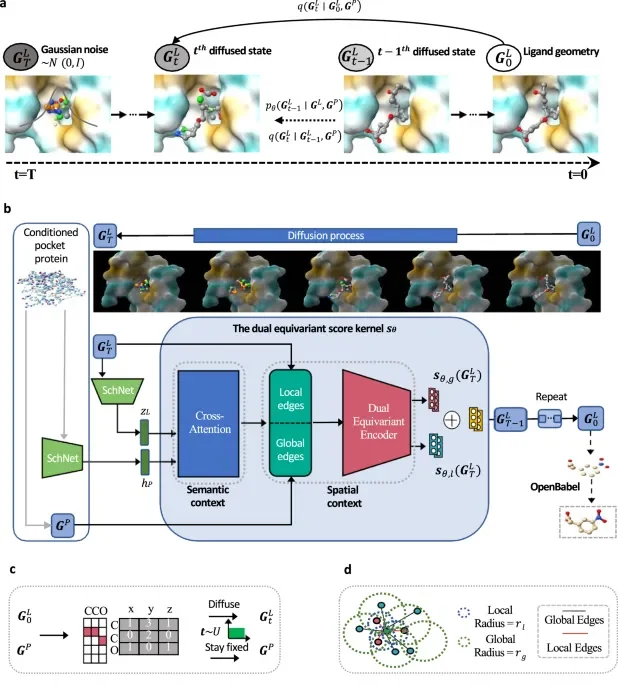
Crafting Molecules Atom by Atom
At the core of this revolution are diffusion models. These complex AI architectures, inspired by physical processes of spreading and refining, learn to generate intricate molecular structures.
Read More: New Brain Scans May Show Vegetative State Patients Are Aware
They can design novel compounds with specific characteristics, such as solubility or size, crucial for a drug's effectiveness and delivery.
Techniques like 'AI-augmented molecular docking' and 'end-to-end binding affinity prediction' are being integrated, allowing for more accurate assessments of how well a potential drug will bind to its intended target.
Some systems are even incorporating 'quantum chemistry-informed neural potentials', bringing deeper physical understanding into the design process.
Tailored Therapies for Specific Ailments
The potential extends beyond general drug discovery. These AI tools are being honed for highly specific applications:

In cancer research, models are being developed to generate anti-cancer drugs designed to combat specific genetic profiles of tumors, aiming to bypass resistance mechanisms. This involves predicting how compounds will interact with both sensitive and resistant cancer cells.
Systems are also focusing on generating 3D molecular structures directly, aligning them with the 'pockets' of target proteins. This geometric precision is vital for effective drug binding.
The Digital Laboratory Takes Shape
This paradigm shift leverages diverse AI techniques. Beyond diffusion models, other generative AI approaches are at play:
'Variational autoencoders' (VAEs) and 'generative adversarial networks' (GANs) are established tools for exploring chemical spaces.
'Autoregressive transformers' and 'Large Language Models' (LLMs) are being adapted for modeling protein sequences and guiding molecular design.
The ultimate vision is the synthesis of these AI capabilities with automated laboratory equipment and even quantum computing, creating self-directed 'molecular design ecosystems'.
The rapid development in this field is evident in the publication dates of recent research. Significant work on diffusion models for drug design appeared throughout 2025, with foundational techniques emerging in 2024.
Notable Developments
DrugDiff: A diffusion model designed to generate small molecules with a range of specified properties, demonstrated by creating thousands of molecules for different scenarios.
Genotype-to-drug models: AI systems specifically trained to design cancer therapeutics based on a patient's genetic information.
Dual diffusion models: Tools that simultaneously generate 3D molecules and refine them for optimal fit within protein targets.
DiffDock and RFdiffusion: Examples of AI platforms that have shown success in protein engineering and molecular docking.








