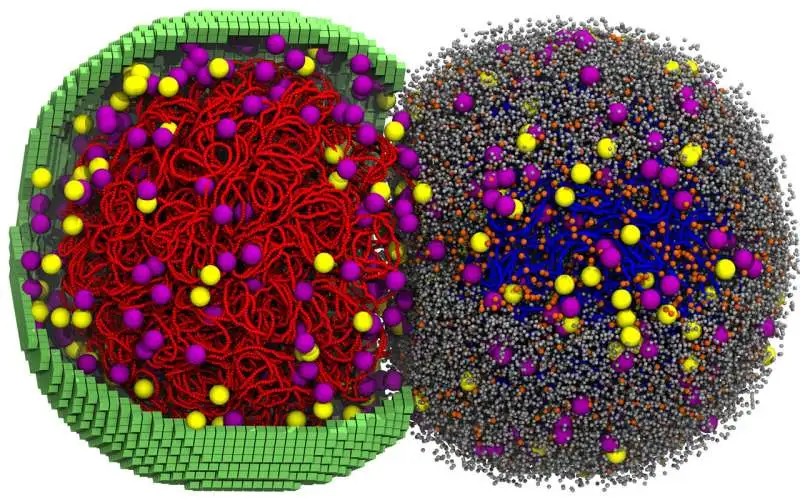
The attempt to turn life into a computational mirror has reached a state of crude mimicry. Researchers have finished a simulation of a living cell across its entire cycle, mapping the jitter and collision of every molecule at a nanoscale resolution. This digital double tracked a 105-minute life span, requiring six days of continuous GPU crunching to reach its conclusion.
The simulation finished within two minutes of actual biological timing.
The model accounted for the physical swelling required before a cell splits.
Every internal molecule was accounted for, from energy consumption to structural drift.
Calculating the Drag of Existence
This wasn't a simulation of a complex human cell, which carries a messy burden of 20,000 genes, but a minimal cell—a laboratory artifact stripped of everything but the bare essentials for survival. By stripping the organism down to its pared-down genetic kit, the team created a closed system that the machines could finally digest.
"The ability to accurately capture the ever-changing conditions within a living cell opens a new window on the foundations of living systems." — Luthey-Schulten
| Metric | Simulated Life | Physical Life |
|---|---|---|
| Duration | 6 Days (GPU time) | 105 Minutes |
| Resolution | Nanoscale | Molecular |
| Accuracy Gap | ~2 Minutes | N/A |
| Gene Count | Minimal Set | ~5,000 (E. coli) |
The data suggests a lopsided economy of scale within the organism. The minimal cell is a parasite, and its primary labor is not creation, but transporting molecules across its outer skin. Most of its chemical currency is spent simply moving things from the outside to the inside, a relentless mechanical chore that defines its existence.
Read More: arXiv Bans Authors for Unchecked AI Content for One Year
Hardware as the New Petri Dish
The bottleneck for this work was not just the code, but the raw heat and math provided by NVIDIA GPUs. To get a single "run" to work, the team had to wait nearly a week for the hardware to iterate through the billions of tiny collisions.
Multiple GPUs worked in parallel to resolve the nanoscale resolution.
Start conditions were slightly tweaked in repeated tests to ensure the cell didn't just survive by luck of the first digit.
The process highlighted that even a "simple" cell is a data-heavy burden that nearly chokes modern processors.
Background: The Reduced Organism
The project, published in the journal Cell, relies on the concept of the "minimal cell." Unlike an E. coli cell, which manages roughly 5,000 genes, the subject of this study is a biological skeleton. This research is less about the "soul" of life and more about the logistics of a container. By simulating the doubling of size and the eventual snap of division, the researchers are trying to see if biology is just a very complex set of rules that can be translated into silicon without losing the essence of the "real" thing. For now, the "real" thing is 80 times faster than its digital ghost.